DigiSkin Atlas
An open resource for RNA-sequencing data of the skin.
RNA sequence data provide insight into which genes are currently being transcribed and utilized in the skin. This enables us to characterize the phenotype and behavior of cells. Currently, sequencing data has to be generated anew in each laboratory because data from previously published studies is fragmentary and often inaccessible to other researchers. We want to change that.
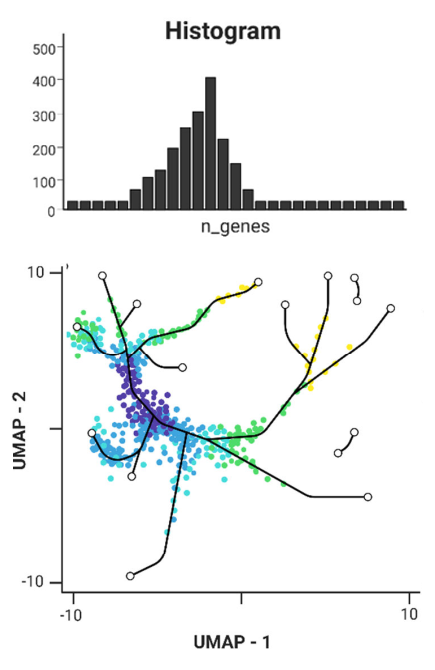
A robust data set has already been created from 20 published studies using single cell sequencing data from mouse skin. We provide proof-of-concept evidence that the dataset enables detection of small populations of cells such as senescent cells. Moreover, we verify the consistency of the in silico analysis with already published results and in vivo analysis.
Our goal now is to expand this platform by integrating more data from different genotypes, skin diseases and skin appendages. In collaboration with experts, a digital skin atlas will be created that will not only reduce the need for animal models, but also provide groundbreaking insights into skin biology and cellular senescence. The DigiSkin Atlas will be made available online to provide scientists with a powerful resource for researching various diseases and phenotypes in mouse skin.
In our lab, we will use the dataset to analyse the features of cellular senescence in relation to the factors related to wound healing such as epithelial-to-mesenchymal transition markers, migration-promoting factors, cytokines, and others. This will be done in skin cell types including keratinocytes, fibroblasts, immune cells, and verified by in situ histological experiments.
The project is made possible via a 3R/Stand-Alone grant by FWF (Grant-DOI: 10.55776/P37321). Thank you for your support!